Estimating effect of multiple treatments#
[1]:
import dowhy
dowhy.enable_notebook_rendering()
from dowhy import CausalModel
import dowhy.datasets
import warnings
warnings.filterwarnings('ignore')
[2]:
data = dowhy.datasets.linear_dataset(10, num_common_causes=4, num_samples=10000,
num_instruments=0, num_effect_modifiers=2,
num_treatments=2,
treatment_is_binary=False,
num_discrete_common_causes=2,
num_discrete_effect_modifiers=0,
one_hot_encode=False)
df=data['df']
df.head()
[2]:
| X0 | X1 | W0 | W1 | W2 | W3 | v0 | v1 | y | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | -1.631834 | -0.732712 | -1.486485 | -0.115370 | 0 | 0 | -8.839621 | -4.979303 | -523.811632 |
| 1 | -1.132617 | 0.715087 | -1.309199 | 0.115198 | 3 | 2 | 8.987890 | 20.536670 | -544.900202 |
| 2 | 1.251704 | -0.438323 | 0.241551 | -0.664516 | 0 | 0 | -2.685975 | -2.140839 | -19.369511 |
| 3 | -0.171360 | 0.979683 | 0.309194 | 0.977418 | 0 | 1 | 10.036483 | 7.047617 | 201.823558 |
| 4 | 1.112863 | 1.672066 | 1.695142 | -0.429582 | 1 | 1 | 10.428849 | 12.888579 | 1214.562517 |
[3]:
model = CausalModel(data=data["df"],
treatment=data["treatment_name"], outcome=data["outcome_name"],
graph=data["gml_graph"])
[4]:
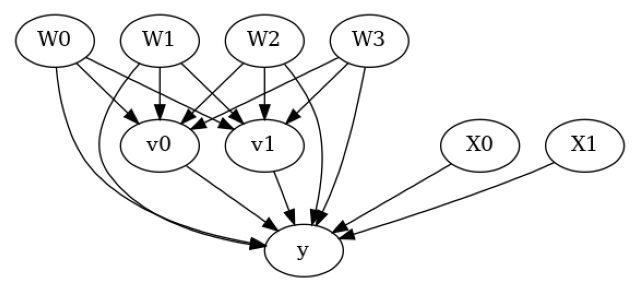
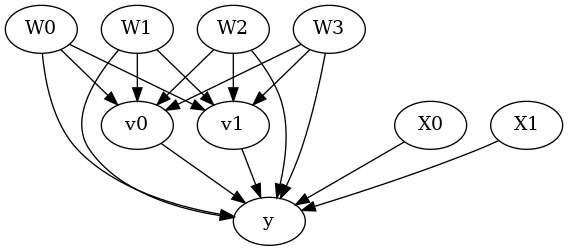
model.view_model()
from IPython.display import Image, display
display(Image(filename="causal_model.png"))
[5]:
identified_estimand= model.identify_effect(proceed_when_unidentifiable=True)
print(identified_estimand)
Estimand type: EstimandType.NONPARAMETRIC_ATE
### Estimand : 1
Estimand name: backdoor
Estimand expression:
d
─────────(E[y|W2,W3,W1,W0])
d[v₀ v₁]
Estimand assumption 1, Unconfoundedness: If U→{v0,v1} and U→y then P(y|v0,v1,W2,W3,W1,W0,U) = P(y|v0,v1,W2,W3,W1,W0)
### Estimand : 2
Estimand name: iv
No such variable(s) found!
### Estimand : 3
Estimand name: frontdoor
No such variable(s) found!
Linear model#
Let us first see an example for a linear model. The control_value and treatment_value can be provided as a tuple/list when the treatment is multi-dimensional.
The interpretation is change in y when v0 and v1 are changed from (0,0) to (1,1).
[6]:
linear_estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression",
control_value=(0,0),
treatment_value=(1,1),
method_params={'need_conditional_estimates': False})
print(linear_estimate)
*** Causal Estimate ***
## Identified estimand
Estimand type: EstimandType.NONPARAMETRIC_ATE
### Estimand : 1
Estimand name: backdoor
Estimand expression:
d
─────────(E[y|W2,W3,W1,W0])
d[v₀ v₁]
Estimand assumption 1, Unconfoundedness: If U→{v0,v1} and U→y then P(y|v0,v1,W2,W3,W1,W0,U) = P(y|v0,v1,W2,W3,W1,W0)
## Realized estimand
b: y~v0+v1+W2+W3+W1+W0+v0*X0+v0*X1+v1*X0+v1*X1
Target units: ate
## Estimate
Mean value: 32.6551274218245
You can estimate conditional effects, based on effect modifiers.
[7]:
linear_estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression",
control_value=(0,0),
treatment_value=(1,1))
print(linear_estimate)
*** Causal Estimate ***
## Identified estimand
Estimand type: EstimandType.NONPARAMETRIC_ATE
### Estimand : 1
Estimand name: backdoor
Estimand expression:
d
─────────(E[y|W2,W3,W1,W0])
d[v₀ v₁]
Estimand assumption 1, Unconfoundedness: If U→{v0,v1} and U→y then P(y|v0,v1,W2,W3,W1,W0,U) = P(y|v0,v1,W2,W3,W1,W0)
## Realized estimand
b: y~v0+v1+W2+W3+W1+W0+v0*X0+v0*X1+v1*X0+v1*X1
Target units:
## Estimate
Mean value: 32.6551274218245
### Conditional Estimates
__categorical__X0 __categorical__X1
(-3.385, -0.862] (-3.529, -0.329] -125.149971
(-0.329, 0.273] -106.592181
(0.273, 0.765] -95.176969
(0.765, 1.363] -83.803876
(1.363, 4.247] -64.066238
(-0.862, -0.27] (-3.529, -0.329] -46.822087
(-0.329, 0.273] -27.579335
(0.273, 0.765] -17.297589
(0.765, 1.363] -5.062335
(1.363, 4.247] 14.255666
(-0.27, 0.23] (-3.529, -0.329] 2.640733
(-0.329, 0.273] 21.474724
(0.273, 0.765] 32.596247
(0.765, 1.363] 44.018360
(1.363, 4.247] 63.075617
(0.23, 0.808] (-3.529, -0.329] 51.508618
(-0.329, 0.273] 70.768601
(0.273, 0.765] 80.177867
(0.765, 1.363] 92.897428
(1.363, 4.247] 111.034967
(0.808, 4.199] (-3.529, -0.329] 134.206907
(-0.329, 0.273] 149.658092
(0.273, 0.765] 159.859578
(0.765, 1.363] 172.246050
(1.363, 4.247] 187.855341
dtype: float64
More methods#
You can also use methods from EconML or CausalML libraries that support multiple treatments. You can look at examples from the conditional effect notebook: https://py-why.github.io/dowhy/example_notebooks/dowhy-conditional-treatment-effects.html
Propensity-based methods do not support multiple treatments currently.