Demo for the DoWhy causal API#
We show a simple example of adding a causal extension to any dataframe.
[1]:
import dowhy
dowhy.enable_notebook_rendering()
import dowhy.datasets
import dowhy.api
from dowhy.graph import build_graph_from_str
import numpy as np
import pandas as pd
from statsmodels.api import OLS
[2]:
data = dowhy.datasets.linear_dataset(beta=5,
num_common_causes=1,
num_instruments = 0,
num_samples=1000,
treatment_is_binary=True)
df = data['df']
df['y'] = df['y'] + np.random.normal(size=len(df)) # Adding noise to data. Without noise, the variance in Y|X, Z is zero, and mcmc fails.
nx_graph = build_graph_from_str(data["dot_graph"])
treatment= data["treatment_name"][0]
outcome = data["outcome_name"][0]
common_cause = data["common_causes_names"][0]
df
[2]:
| W0 | v0 | y | |
|---|---|---|---|
| 0 | -0.100021 | False | 0.132437 |
| 1 | 0.534403 | True | 4.776436 |
| 2 | 0.111168 | False | -0.824853 |
| 3 | 1.370331 | True | 8.595711 |
| 4 | -0.618126 | False | -0.541416 |
| ... | ... | ... | ... |
| 995 | -0.379531 | True | 2.793668 |
| 996 | -1.785375 | True | 2.363306 |
| 997 | -2.354834 | True | 1.268439 |
| 998 | -0.957736 | True | 3.688295 |
| 999 | -1.240677 | False | -0.713537 |
1000 rows × 3 columns
[3]:
# data['df'] is just a regular pandas.DataFrame
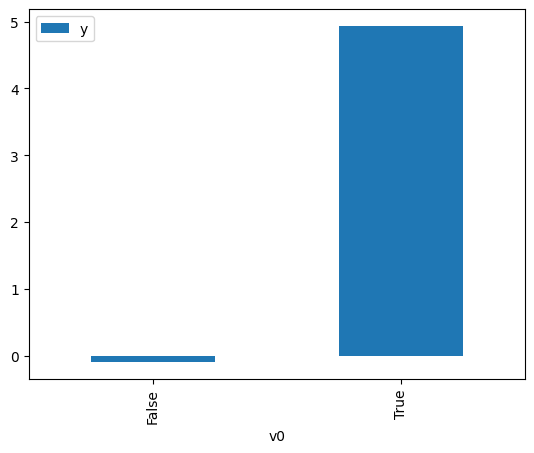
df.causal.do(x=treatment,
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
common_causes=[common_cause],
).groupby(treatment).mean().plot(y=outcome, kind='bar')
[3]:
<Axes: xlabel='v0'>
[4]:
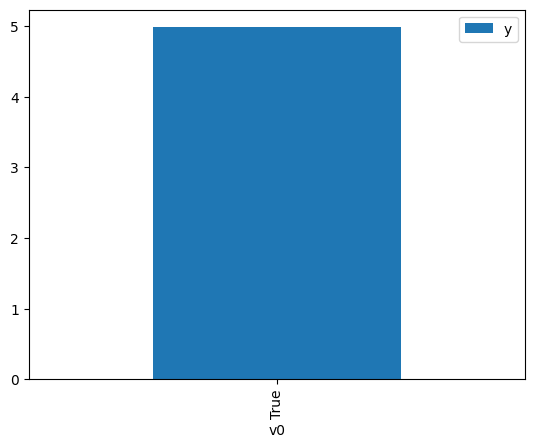
df.causal.do(x={treatment: 1},
variable_types={treatment:'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
method='weighting',
common_causes=[common_cause]
).groupby(treatment).mean().plot(y=outcome, kind='bar')
[4]:
<Axes: xlabel='v0'>
[5]:
cdf_1 = df.causal.do(x={treatment: 1},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
graph=nx_graph
)
cdf_0 = df.causal.do(x={treatment: 0},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
graph=nx_graph
)
[6]:
cdf_0
[6]:
| W0 | v0 | y | propensity_score | weight | |
|---|---|---|---|---|---|
| 0 | -0.279066 | False | 0.311358 | 0.504760 | 1.981141 |
| 1 | -0.355988 | False | -0.875442 | 0.507923 | 1.968801 |
| 2 | 1.127934 | False | 2.253372 | 0.447079 | 2.236739 |
| 3 | 0.929628 | False | 1.149448 | 0.455159 | 2.197035 |
| 4 | 1.048707 | False | 2.814462 | 0.450304 | 2.220721 |
| ... | ... | ... | ... | ... | ... |
| 995 | 1.967333 | False | 3.934154 | 0.413237 | 2.419917 |
| 996 | -0.929117 | False | -2.203187 | 0.531459 | 1.881612 |
| 997 | -1.862456 | False | -2.065187 | 0.569443 | 1.756101 |
| 998 | -0.014084 | False | 0.942722 | 0.493860 | 2.024867 |
| 999 | 0.373045 | False | 0.639678 | 0.477948 | 2.092276 |
1000 rows × 5 columns
[7]:
cdf_1
[7]:
| W0 | v0 | y | propensity_score | weight | |
|---|---|---|---|---|---|
| 0 | -1.016380 | True | 2.974043 | 0.464967 | 2.150690 |
| 1 | -1.750903 | True | 3.555488 | 0.435063 | 2.298520 |
| 2 | -0.605960 | True | 2.955557 | 0.481801 | 2.075546 |
| 3 | -0.125322 | True | 4.851993 | 0.501565 | 1.993761 |
| 4 | -1.366810 | True | 2.508822 | 0.450655 | 2.218991 |
| ... | ... | ... | ... | ... | ... |
| 995 | 0.318000 | True | 8.975365 | 0.519791 | 1.923849 |
| 996 | 1.241249 | True | 5.593692 | 0.557525 | 1.793641 |
| 997 | -1.381995 | True | 3.793374 | 0.450037 | 2.222041 |
| 998 | 0.801727 | True | 7.344791 | 0.539617 | 1.853165 |
| 999 | -1.030269 | True | 3.333833 | 0.464399 | 2.153323 |
1000 rows × 5 columns
Comparing the estimate to Linear Regression#
First, estimating the effect using the causal data frame, and the 95% confidence interval.
[8]:
(cdf_1['y'] - cdf_0['y']).mean()
[8]:
$\displaystyle 5.06699896618239$
[9]:
1.96*(cdf_1['y'] - cdf_0['y']).std() / np.sqrt(len(df))
[9]:
$\displaystyle 0.163468403376322$
Comparing to the estimate from OLS.
[10]:
model = OLS(np.asarray(df[outcome]), np.asarray(df[[common_cause, treatment]], dtype=np.float64))
result = model.fit()
result.summary()
[10]:
| Dep. Variable: | y | R-squared (uncentered): | 0.935 |
|---|---|---|---|
| Model: | OLS | Adj. R-squared (uncentered): | 0.935 |
| Method: | Least Squares | F-statistic: | 7149. |
| Date: | Wed, 22 Apr 2026 | Prob (F-statistic): | 0.00 |
| Time: | 08:22:17 | Log-Likelihood: | -1405.6 |
| No. Observations: | 1000 | AIC: | 2815. |
| Df Residuals: | 998 | BIC: | 2825. |
| Df Model: | 2 | ||
| Covariance Type: | nonrobust |
| coef | std err | t | P>|t| | [0.025 | 0.975] | |
|---|---|---|---|---|---|---|
| x1 | 1.6382 | 0.030 | 54.469 | 0.000 | 1.579 | 1.697 |
| x2 | 5.0559 | 0.045 | 112.840 | 0.000 | 4.968 | 5.144 |
| Omnibus: | 0.105 | Durbin-Watson: | 1.958 |
|---|---|---|---|
| Prob(Omnibus): | 0.949 | Jarque-Bera (JB): | 0.176 |
| Skew: | -0.005 | Prob(JB): | 0.916 |
| Kurtosis: | 2.936 | Cond. No. | 1.53 |
Notes:
[1] R² is computed without centering (uncentered) since the model does not contain a constant.
[2] Standard Errors assume that the covariance matrix of the errors is correctly specified.